1. Basic Information
| Name: | PK3CD_HUMAN |
| Accession#: | O00329 |
| Description: | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta isoform |
| AA Number: | 1044 |
| Sequence: |
1
51
101
151
201
251
301
351
401
451
501
551
601
651
701
751
801
851
901
951
1001
|
|
MPPGVDCPME FWTKEENQSV VVDFLLPTGV YLNFPVSRNA NLSTIKQLLW
HRAQYEPLFH MLSGPEAYVF TCINQTAEQQ ELEDEQRRLC DVQPFLPVLR
LVAREGDRVK KLINSQISLL IGKGLHEFDS LCDPEVNDFR AKMCQFCEEA
AARRQQLGWE AWLQYSFPLQ LEPSAQTWGP GTLRLPNRAL LVNVKFEGSE
ESFTFQVSTK DVPLALMACA LRKKATVFRQ PLVEQPEDYT LQVNGRHEYL
YGSYPLCQFQ YICSCLHSGL TPHLTMVHSS SILAMRDEQS NPAPQVQKPR
AKPPPIPAKK PSSVSLWSLE QPFRIELIQG SKVNADERMK LVVQAGLFHG
NEMLCKTVSS SEVSVCSEPV WKQRLEFDIN ICDLPRMARL CFALYAVIEK
AKKARSTKKK SKKADCPIAW ANLMLFDYKD QLKTGERCLY MWPSVPDEKG
ELLNPTGTVR SNPNTDSAAA LLICLPEVAP HPVYYPALEK ILELGRHSEC
VHVTEEEQLQ LREILERRGS GELYEHEKDL VWKLRHEVQE HFPEALARLL
LVTKWNKHED VAQMLYLLCS WPELPVLSAL ELLDFSFPDC HVGSFAIKSL
RKLTDDELFQ YLLQLVQVLK YESYLDCELT KFLLDRALAN RKIGHFLFWH
LRSEMHVPSV ALRFGLILEA YCRGSTHHMK VLMKQGEALS KLKALNDFVK
LSSQKTPKPQ TKELMHLCMR QEAYLEALSH LQSPLDPSTL LAEVCVEQCT
FMDSKMKPLW IMYSNEEAGS GGSVGIIFKN GDDLRQDMLT LQMIQLMDVL
WKQEGLDLRM TPYGCLPTGD RTGLIEVVLR SDTIANIQLN KSNMAATAAF
NKDALLNWLK SKNPGEALDR AIEEFTLSCA GYCVATYVLG IGDRHSDNIM
IRESGQLFHI DFGHFLGNFK TKFGINRERV PFILTYDFVH VIQQGKTNNS
EKFERFRGYC ERAYTILRRH GLLFLHLFAL MRAAGLPELS CSKDIQYLKD
SLALGKTEEE ALKHFRVKFN EALRESWKTK VNWLAHNVSK DNRQ |
|
*Highlighted peptides (with yellow background) have developed assays.
*Green background amino acids are PTMs.
2. Protein Separation
Sample Preparation: Cells from colon caner line HCT 116 were lysed in 8M urea/ 100mM ammonium bicarbonate buffer on ice. The equivalent of 60,000 cells were loaded onto a SDS gel. The protein of interest was excised and ingel digestion with trypsin was performed after reduction and alkylation with TCEP and IAA. 1/6 of the resulting digest was then analyzed on a Thermo Scientific TSQ mass spectrometer.
3. LC-MS/MS Data
GELLNPTGTVR
No LC-MS/MS for this peptide.
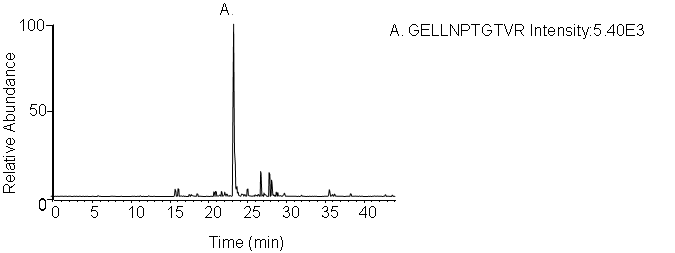
4. LC-MRM Screening
Peptides screening: Peptides and transitions for analysis were selected using a predictive peptide program. The results were screened for those with the highest specificity and the most intense transitions.