1. Basic Information
| Name: | KI67_HUMAN |
| Accession#: | P46013 |
| Description: | Antigen KI-67 |
| AA Number: | 3256 |
| Sequence: |
1
51
101
151
201
251
301
351
401
451
501
551
601
651
701
751
801
851
901
951
1001
1051
1101
1151
1201
1251
1301
1351
1401
1451
1501
1551
1601
1651
1701
1751
1801
1851
1901
1951
2001
2051
2101
2151
2201
2251
2301
2351
2401
2451
2501
2551
2601
2651
2701
2751
2801
2851
2901
2951
3001
3051
3101
3151
3201
3251
|
|
MWPTRRLVTI KRSGVDGPHF PLSLSTCLFG RGIECDIRIQ LPVVSKQHCK
IEIHEQEAIL HNFSSTNPTQ VNGSVIDEPV RLKHGDVITI IDRSFRYENE
SLQNGRKSTE FPRKIREQEP ARRVSRSSFS SDPDEKAQDS KAYSKITEGK
VSGNPQVHIK NVKEDSTADD SKDSVAQGTT NVHSSEHAGR NGRNAADPIS
GDFKEISSVK LVSRYGELKS VPTTQCLDNS KKNESPFWKL YESVKKELDV
KSQKENVLQY CRKSGLQTDY ATEKESADGL QGETQLLVSR KSRPKSGGSG
HAVAEPASPE QELDQNKGKG RDVESVQTPS KAVGASFPLY EPAKMKTPVQ
YSQQQNSPQK HKNKDLYTTG RRESVNLGKS EGFKAGDKTL TPRKLSTRNR
TPAKVEDAAD SATKPENLSS KTRGSIPTDV EVLPTETEIH NEPFLTLWLT
QVERKIQKDS LSKPEKLGTT AGQMCSGLPG LSSVDINNFG DSINESEGIP
LKRRRVSFGG HLRPELFDEN LPPNTPLKRG EAPTKRKSLV MHTPPVLKKI
IKEQPQPSGK QESGSEIHVE VKAQSLVISP PAPSPRKTPV ASDQRRRSCK
TAPASSSKSQ TEVPKRGGRK SGNLPSKRVS ISRSQHDILQ MICSKRRSGA
SEANLIVAKS WADVVKLGAK QTQTKVIKHG PQRSMNKRQR RPATPKKPVG
EVHSQFSTGH ANSPCTIIIG KAHTEKVHVP ARPYRVLNNF ISNQKMDFKE
DLSGIAEMFK TPVKEQPQLT STCHIAISNS ENLLGKQFQG TDSGEEPLLP
TSESFGGNVF FSAQNAAKQP SDKCSASPPL RRQCIRENGN VAKTPRNTYK
MTSLETKTSD TETEPSKTVS TANRSGRSTE FRNIQKLPVE SKSEETNTEI
VECILKRGQK ATLLQQRREG EMKEIERPFE TYKENIELKE NDEKMKAMKR
SRTWGQKCAP MSDLTDLKSL PDTELMKDTA RGQNLLQTQD HAKAPKSEKG
KITKMPCQSL QPEPINTPTH TKQQLKASLG KVGVKEELLA VGKFTRTSGE
TTHTHREPAG DGKSIRTFKE SPKQILDPAA RVTGMKKWPR TPKEEAQSLE
DLAGFKELFQ TPGPSEESMT DEKTTKIACK SPPPESVDTP TSTKQWPKRS
LRKADVEEEF LALRKLTPSA GKAMLTPKPA GGDEKDIKAF MGTPVQKLDL
AGTLPGSKRQ LQTPKEKAQA LEDLAGFKEL FQTPGHTEEL VAAGKTTKIP
CDSPQSDPVD TPTSTKQRPK RSIRKADVEG ELLACRNLMP SAGKAMHTPK
PSVGEEKDII IFVGTPVQKL DLTENLTGSK RRPQTPKEEA QALEDLTGFK
ELFQTPGHTE EAVAAGKTTK MPCESSPPES ADTPTSTRRQ PKTPLEKRDV
QKELSALKKL TQTSGETTHT DKVPGGEDKS INAFRETAKQ KLDPAASVTG
SKRHPKTKEK AQPLEDLAGL KELFQTPVCT DKPTTHEKTT KIACRSQPDP
VDTPTSSKPQ SKRSLRKVDV EEEFFALRKR TPSAGKAMHT PKPAVSGEKN
IYAFMGTPVQ KLDLTENLTG SKRRLQTPKE KAQALEDLAG FKELFQTRGH
TEESMTNDKT AKVACKSSQP DPDKNPASSK RRLKTSLGKV GVKEELLAVG
KLTQTSGETT HTHTEPTGDG KSMKAFMESP KQILDSAASL TGSKRQLRTP
KGKSEVPEDL AGFIELFQTP SHTKESMTNE KTTKVSYRAS QPDLVDTPTS
SKPQPKRSLR KADTEEEFLA FRKQTPSAGK AMHTPKPAVG EEKDINTFLG
TPVQKLDQPG NLPGSNRRLQ TRKEKAQALE ELTGFRELFQ TPCTDNPTTD
EKTTKKILCK SPQSDPADTP TNTKQRPKRS LKKADVEEEF LAFRKLTPSA
GKAMHTPKAA VGEEKDINTF VGTPVEKLDL LGNLPGSKRR PQTPKEKAKA
LEDLAGFKEL FQTPGHTEES MTDDKITEVS CKSPQPDPVK TPTSSKQRLK
ISLGKVGVKE EVLPVGKLTQ TSGKTTQTHR ETAGDGKSIK AFKESAKQML
DPANYGTGME RWPRTPKEEA QSLEDLAGFK ELFQTPDHTE ESTTDDKTTK
IACKSPPPES MDTPTSTRRR PKTPLGKRDI VEELSALKQL TQTTHTDKVP
GDEDKGINVF RETAKQKLDP AASVTGSKRQ PRTPKGKAQP LEDLAGLKEL
FQTPICTDKP TTHEKTTKIA CRSPQPDPVG TPTIFKPQSK RSLRKADVEE
ESLALRKRTP SVGKAMDTPK PAGGDEKDMK AFMGTPVQKL DLPGNLPGSK
RWPQTPKEKA QALEDLAGFK ELFQTPGTDK PTTDEKTTKI ACKSPQPDPV
DTPASTKQRP KRNLRKADVE EEFLALRKRT PSAGKAMDTP KPAVSDEKNI
NTFVETPVQK LDLLGNLPGS KRQPQTPKEK AEALEDLVGF KELFQTPGHT
EESMTDDKIT EVSCKSPQPE SFKTSRSSKQ RLKIPLVKVD MKEEPLAVSK
LTRTSGETTQ THTEPTGDSK SIKAFKESPK QILDPAASVT GSRRQLRTRK
EKARALEDLV DFKELFSAPG HTEESMTIDK NTKIPCKSPP PELTDTATST
KRCPKTRPRK EVKEELSAVE RLTQTSGQST HTHKEPASGD EGIKVLKQRA
KKKPNPVEEE PSRRRPRAPK EKAQPLEDLA GFTELSETSG HTQESLTAGK
ATKIPCESPP LEVVDTTAST KRHLRTRVQK VQVKEEPSAV KFTQTSGETT
DADKEPAGED KGIKALKESA KQTPAPAASV TGSRRRPRAP RESAQAIEDL
AGFKDPAAGH TEESMTDDKT TKIPCKSSPE LEDTATSSKR RPRTRAQKVE
VKEELLAVGK LTQTSGETTH TDKEPVGEGK GTKAFKQPAK RKLDAEDVIG
SRRQPRAPKE KAQPLEDLAS FQELSQTPGH TEELANGAAD SFTSAPKQTP
DSGKPLKISR RVLRAPKVEP VGDVVSTRDP VKSQSKSNTS LPPLPFKRGG
GKDGSVTGTK RLRCMPAPEE IVEELPASKK QRVAPRARGK SSEPVVIMKR
SLRTSAKRIE PAEELNSNDM KTNKEEHKLQ DSVPENKGIS LRSRRQNKTE
AEQQITEVFV LAERIEINRN EKKPMKTSPE MDIQNPDDGA RKPIPRDKVT
ENKRCLRSAR QNESSQPKVA EESGGQKSAK VLMQNQKGKG EAGNSDSMCL
RSRKTKSQPA ASTLESKSVQ RVTRSVKRCA ENPKKAEDNV CVKKIRTRSH
RDSEDI |
|
*Highlighted peptides (with yellow background) have developed assays.
*Green background amino acids are PTMs.
2. Protein Separation
Sample Preparation: Cells from Breast cancer MCF7 cell line were lysed in 8M urea/ 100mM ammonium bicarbonate buffer on ice. The equivalent of 100,000 cells were loaded onto a SDS gel. The protein of interest was excised and ingel digestion with trypsin was performed after reduction and alkylation with TCEP and IAA. 1/4 of the resulting digest was then analyzed on a Thermo Scientific TSQ mass spectrometer.
3. LC-MS/MS Data
SPPPESVDTPTSTK
No LC-MS/MS for this peptide.
VHVPARPYR
No LC-MS/MS for this peptide.
TPVQYSQQQNSPQK
No LC-MS/MS for this peptide.
DSLSKPEK
No LC-MS/MS for this peptide.
TSDTETEPSK
No LC-MS/MS for this peptide.
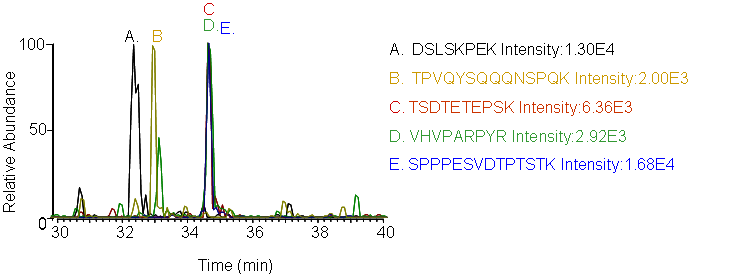
4. LC-MRM Screening
Peptides screening: Peptides and transitions for analysis were selected using a predictive peptide program. The results were screened for those with the highest specificity and the most intense transitions.